
The ruminant exploits a digestive strategy that relies on an intimate relationship with various microorganisms and, as such, can utilize feedstuffs that other animals cannot.
Mammals have little to no ability to degrade fibrous plant materials, but bacteria, protozoa and fungi produce cellulases, a group of enzymes able to breakdown cellulose and other related polysaccharides found in plant cell wall material. The ruminant foregut houses an ecosystem of these anaerobic microbes that allow ruminants to obtain nutrients from not only fibrous plant-based materials, but also the microbes themselves — meaning ruminant agriculture can produce human-edible products from human-inedible forage.
Rumen fermentation
The rumen essentially acts as a large fermentation vessel where bacteria, protozoa and fungi digest and degrade fibrous and non-fibrous plant material, as well as protein and non-protein nitrogen sources. There is also digestion and modification of lipids by bacterial lipases and biohydrogenation.
End products of rumen fermentation are either absorbed directly through the rumen wall, for example volatile fatty acids (VFA), or in the small intestine (SI), for example small peptides/amino acids and lipids. Dead microbes are also washed through the forestomach and can be digested and absorbed in the SI and provide a significant contribution to the host’s nitrogen requirement.
The rumen ecosystem is made up of four groups of microbes: bacteria, protozoa, fungi and archaea. There are also bacteriophages, mycoplasmas and archaeophages. In terms of numbers, bacteria are by far the most abundant at 1010 – 1011cells/ml, followed by archaea (106 – 108 cells/ml), protozoa (105 cells/ml) and fungi (103 – 105 cells/ml). Bacteria comprise about two-thirds of the rumen microbial biomass with protozoa making up to 50 percent due mainly to their size.
Examination of rumen bacteria
Since the 1980s, modern molecular techniques have allowed us to better understand the role of the microbial ecosystem and particularly the diversity within it. The 16S rRNA gene was a breakthrough in terms of being able to catalogue changes in bacterial communities because of different rumen environments, including diets. Pyrosequencing of a specific region of the 16S rRNA gene allows taxonomic classification of rumen bacteria and the ability to evaluate bacteria diversity in the rumen.
While much research has historically focused on individual strains, recent research has started to look at the bacterial community as a whole in response to changes in rumen environment, namely dietary changes.
Bacteria are a diverse, plastic group with a core community that appears relatively consistent across diets. Studies of rumen bacteria have shown three major phyla that comprise the core community, irrespective of diet: Bacteriodetes, Firmicutes and Proteobacteria.
A large-scale study examining microbial communities across a range of ruminant and camelid species, diets and geographical regions found seven bacterial groups accounted for more than 60 percent of the total bacterial samples they sequenced. The authors noted these groups comprised of three known groups (Prevotella, Butyrivibrio, and Ruminococcus), as well as four unclassified groups (Lachnospiraceae, Ruminococcaceae, Bacteroidales, and Clostridiales), but the relative abundance differed for all seven across species. They hypothesized that even though we can identify the “core” species, more research is required in order to understand them and their role in the microbial community.
Similarly, an earlier meta-analysis revealed 19 bacterial phyla present in the rumen microbiome. According to this meta-analysis (Ribosomal Database Project), 71 percent of the bacterial species have been covered, meaning that 30 percent remain unknown.
A core microbiome notwithstanding, diet appears to have a major effect on bacterial diversity.
Across regions, communities from forage-fed animals were similar but distinct from concentrate-fed animals, which were similar to each other. Both were distinct from those fed mixed diets, which were intermediate between the forage-fed and concentrate-fed animals.
Cellulolytic bacteria, such as Bacteroidales, appeared to increase in abundance in those animals fed forage, while propionate producers, such as Prevotella, appeared to increase in the concentrate-fed animals. Similar effects have been seen in other studies.
Rumen protozoa generate interest
Protozoa are comparatively large microbes that are present in the rumen at a concentration of ~105cells/ml. They play a key role in providing nutrients for a host, as well as the generation of methane. Protozoa exhibit a high production of butyrate and acetate, which yields hydrogen molecules that are subsequently taken up and converted to methane by methanogenic archaea. Ruminal protozoa can be sub-divided into two groups: ciliated and flagellated. Despite their other activities, it is the methane production that has received the greatest interest from the scientific community.
It is known that protozoa predate bacteria within the rumen microbial community and this has a negative impact on microbial protein synthesis and, subsequently, nitrogen use efficiency of the cow. A study attempted to elucidate which groups of protozoa were responsible for this predation to suppression of this negative effect. Using labeled rumen bacteria, they discovered that Entodinum sp. were responsible for three-fourths of the predation activity in comparison with holotrich protozoa whose activity was insignificant.
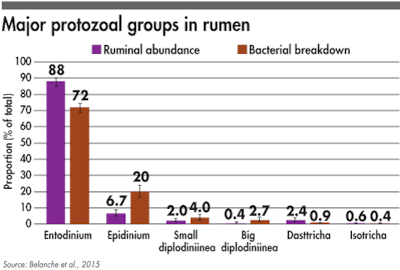
The relative abundance of the six major protozoal groups in rumen of cattle and bacterial breakdown attributed to each of these protozoa groups.
Protozoa have also been implicated in the generation of conjugated linoleic acid (CLA) and in vitro cultures of rumen bacteria, protozoa and a mixture of both revealed that only rumen bacteria were able to fully biohydrogenate linoleic acid to C18:0.
Although many studies exist looking at protozoal effects, and it is well established that defaunation results in a reduction in methane output and has also been shown to improve nutrient use efficiency, far less is known about rumen protozoal communities and diversity compared with bacteria.
Rumen fungi affected by diet
Fungi are the largest of the rumen microbes with the longest generation time. There are six genera: the moncentric Neocallimastix, Piromyces, and Caecomyceas and the polycentric Oprinomyces, Anaeromyces, and Cyllamyces.
They are perhaps best known for their ability to degrade fiber by burrowing into cell-wall material, rendering it more accessible to bacterial enzymes and, ultimately, increasing ruminal fiber. Fungi have also been shown to partially biohydrogenate linoleic acid to t11 C18:1.
Like rumen bacteria, fungal diversity is affected by diet.
Archaea and methane
Many archaea are methanogens and are responsible for generating methane from substrates, such as hydrogen and carbon dioxide.
Compared with bacteria, rumen archaea appear much less diverse. However, an increase in methanogen numbers per se does not necessarily translate into greater methane production as rumen conditions may affect expression of genes in methane production. There are known associations between rumen cellulolytic bacteria and archaea, as well as protozoa and archaea. The link is hydrogen, which archaea use to reduce CO2 to methane.
Protozoa are known hydrogen producers and archaea associate with protozoa to make use of the available hydrogen — methane production is thus used as a hydrogen sink to try to avoid significant drops in rumen pH.
Microbial overview
The rumen microbial ecosystem is diverse and rumen health and function relies on this diversity. Rumen bacteria are probably the most well-studied microbial group, yet it is only in the past few years we are starting to get a picture of the extent of the diversity and nature rumen bacterial strains.
Less is known about protozoa, but removal of protozoa from the rumen environment has led to improvements in rumen efficiency. Fungi are not well-understood but certainly have a critical role to play in making fibrous plant cell wall material accessible to rumen bacteria. Archaea are the group that have become increasingly of interest due to their ability to generate methane utilizing hydrogen from the rumen environment.
Given the current political atmosphere regarding ruminant livestock and reducing greenhouse gas emissions, archaea are starting to be extensively researched using new meta-omics techniques.
References available on request.
















