Micro-array technologies provide opportunities to exponentially enhance the effects of nutrients and diet on animal health and production.
Research and development are key to enhancing our knowledge and understanding of animals. Over the decades, there has been a myriad of different scientific “ages” that have led to significant advances in the knowledge of fundamental aspects of form and function in many species. It appears that the current scientific “age” is the era of the “-omics,” e.g., genomics, proteomics and metabolomics. Work at the molecular level has generated as many questions as it has answers and the idea of genetic programming and alterations of gene expression that have positive physiological consequences are becoming increasingly prevalent.
In this current era of the gene, the substantial effects of nutrition on gene expression are starting to emerge in a science termed nutrigenomics.
Before discussing nutrigenomics, it is important to distinguish between nutrigenomics and nutrigenetics. The latter is focused on how the genetic makeup of an individual coordinates a response to diet.
Nutrigenomics attempts to determine the effects of common dietary ingredients on the activity or expression of whole genome, gaining an insight into how nutritional molecules affect metabolic pathways and homeostasis. Nutrigenomics aims to resolve the interaction between diet and genome and identifies molecular mechanisms of nutrient-related health and disease states.
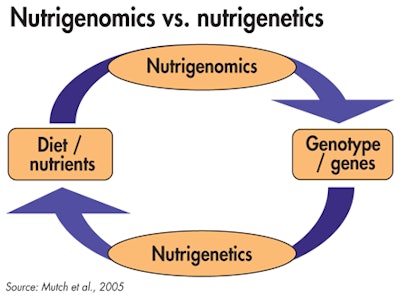 Highlighting the fundamental differences in the approach of nutrigenomics versus nutrigenetics (Mutch et al., 2005)
Highlighting the fundamental differences in the approach of nutrigenomics versus nutrigenetics (Mutch et al., 2005)Exploring gene expression
In recent years, the field has expanded to investigate the impact of classes of nutrients on gene expression and the effect which different chemical forms of the same nutrient have on transcriptional profiles in different tissues.
In human nutrition, hopes are that nutrigenomics will form part of the arsenal of tools to allow the refining of dietary and nutrient recommendations to meet individual requirements. The ability to understand dietary and single nutrient effects on gene expression and, therefore, metabolism will undoubtedly lead to great advances in health, disease and production.
Historically, techniques such as Northern Blots and polymerase chain reaction (PCR) have been used to identify single gene expression, thus requiring knowledge and identification of genes already known to be affected by various endogenous and/or exogenous stimuli. This means sequential, laborious assessment of the expression of individual genes. However, advances in deoxyribonucleic acid (DNA) microarray technology have yielded great results.
The role of microarray technology
Nutrigenomics utilizes DNA microarray technology that has been in use for over a decade. It is capable of profiling gene expression patterns of tens of thousands of genes in a single experiment and it is invaluable when it comes to generating a holistic view of changes in gene expression.
The general construction of microarrays relies on immobilizing DNA sequences — oligonucleotides or amplified cDNA from PCR — onto a solid support, which is then hybridized with a labeled RNA probe. The extent of hybridization of the mRNA with individual sequences on the array gives a quantified measure of gene expression. Genes of specific interest may then undergo sensitive gel-based assays, such as polyacrylamide gel electrophoresis (PAGE).
To fully capitalize on the developing microarray technology application in animals, the fully sequenced genome is required. Otherwise, extrapolation from other species is inevitable, exemplified by the rapid progress in understanding disease in species where the genome has been sequenced.
Elucidation of mechanisms of beneficial dietary components, such as plant sterols and n-3 polyunsaturated fatty acids (PUFA), has been carried out using the technology of microarrays. Identification and quantification of biomarkers for specific disease states have also been aided using this technology and there is a vast potential for other species and diseases.
What microarrays reveal about gene expression
Detailed microarray experiments to investigate the effect of individual nutrients such as selenium have been conducted in animals, including humans, and it demonstrates significant effects on gene expression profiles relating to health and disease.
For example, studies have revealed surprising differences in the gene expression profiles in hen oviduct (a recognized model for the study of reproductive function). These differences resulted from dietary supplementation with inorganic sodium selenite or organic selenium, in the form of selenized yeast, where the organic selenium source affected the expression of key genes involved in the pathway of follicle-stimulating hormone (FSH).
In mice, selenium deficiency resulted in the upregulation of genes associated with stress responses, cell cycling and growth and cell adherence. Concomitantly, there was a downregulation of genes associated with detoxification, selenoprotein production and antioxidant function. These molecular effects can be mapped to the effects seen in selenium-deficient animals.
In humans, work from the Sanders-Brown Center on Aging demonstrated that organic selenium was able to reduce the amyloid plaque burden — a key feature in Alzheimer’s — and minimize DNA and RNA oxidation. Microarray techniques can now look at the exact changes in profiles of gene expression in the pathology of Alzheimer’s to identify the nature of the beneficial effect.
Microarray technology was used to study the effects of fungal infection in grass on luteal function in heifers. The expected clinical effects from fescue intoxication were reflected in the gene expression results, helping the study to demonstrate that gene expression profiling can be used to assess dietary management regimes at the molecular level.
With the development and refining of microarray technologies, the opportunities for the quantification and qualification of the effects of nutrients and diet on health and production, both on an individual, as well as herd level, appear exponential. By determining physiological effects of specific nutrients, it may be possible to improve performance through dietary manipulation.